Discussion, requests for help, suggestions and complaints…
Proton tunneling through graphene (6 replies and 1 comment)
Hello!
A bit more detailed description of your calculations would help others to understand your problem, I think. What do you mean by "not getting physical results"?
Also keep in mind, that you have a charged system for the proton tunneling. In the CRYSTAL manual it is stated, that charged reference cells cannot be used for total energy comparison (see the descrption for the keyword CHARGED).
Hello hanros,
I agree with Christoph Lechner that we need more information in order to help you, and possibly (depending on the specifics) also a copy of your input and output files.
I just wanted to clarify a bit the point of charged cells: since the electrostatic energy of a periodic charged system diverges, it is standard to add an uniform charge background to obtain charge neutrality. This is what the keyword CHARGED does. Since you have a nonphysical interaction with the background, you cannot compare directly the energy with that of a neutral cell (and you cannot compare the energy of cells of different volume). Fortunately there are ways to correct this (for ex., Makov-Payne correction for charged defects) and if you are computing energy differences between different positions of the proton but keeping constant charge and unit cell, the values should be correct. Charged systems can be tricky for other reasons, though, and especially so if they are periodic in 1D or 2D.
By the way, in case you have other problems with CRYSTAL in the future you can also ask support through the online form http://www.crystalsolutions.eu/support.html.
Hi guys!I really wasn't expecting any response, so this is a pleasant surprise. to recap I am trying to tunnel a proton through a benzene ring in a one atom thick graphene sheet. I was able to perform ab initio calculations using crystal14 having hydrogen as the tunneling species but upon removing the shell charges to zero (to simulate a proton), i still get the same results. To clarify i am seeking to attain the barrier energy for proton tunneling by physically changing the vertical location of the proton above the graphene sheet and evaluating the total energy at each point. I know there are more accurate methods such as CINEB and the alternative provided in crystal14 ,reaction co-ordinate method, but both of these methods are unusable at my initial starting position of 2.86 Angstroms above the graphene sheet. I am not using CHARGED cells.I have analysed the atomic charges during the SCF cycle and it seem there is a transfer of electrons from the graphene sheet to the proton, which is entirely unphysical. I have read that this is attributed to DFT global optimization scheme which means the flow of electrons cannot be controlled. i also read that this is due to the breakdown of the Born Oppenheimer approximation in DFT which results in the proton being treated as a point charge.I was wondering whether any of you would be kind enough to advise me on how i could get past this problem.
Thank you, now I think I get the picture.
There is just one thing which is not clear to me. If you are using a periodic cell (as opposed to a "cluster" model) for graphene and you are not using CHARGED, the program should stop if the number of electrons is different from the total nuclear charge. Is it possible that you removed the electron from the hydrogen but you put it somewhere else? If that is the case it is reasonable that the electron will go back to the hydrogen.
I can't find any guidelines of the forum, but if you attached an input file to the next message it might be useful. It is possible that this is a simple input mistake.
Hi Jbaima
I have attached my input file. In this file i have defined a 2x1 supercell, and used ATOMINSE to insert a hydrogen atom at what would be the centre of each benzene ring. As explained above to calculate the barrier energy I calculated the difference in energy between the H atom at 0.0 angstrom and 2.86 Angstrom (physisoprtion state) along the z axis. this file is for the 0.0 Angstrom.i have optimised the geometry of the atomic positions of the carbon atoms only and have kept the z co-ordinates of the carbon atoms fixed. this is on the basis of a recommendation from my professor.To change the charge on the hydrogen atom i have used CHEMOD. i surprisingly got the same energy values with and without CHARGED. i therefore have not used CHARGED in this file.I don't think the use of ATOMSPIN makes any difference as there should be no electron on the hydrogen after use of CHEMOD. ATOMSPIN allows for automatic UHF calculations.I have used NOSHIFT as I believed the fermi level kept shifting upwards into the conduction band during optimisation. this was supported by a professor.for the convergence criteria i have used FMIXING 90 and SMEAR 0.01, as my files previously took too long to converge.Thanks for the help.
Hi Jbaima
I have attached an AVOGADRO image to clarify the positions of the atoms.
Thanks
Hi Jbaima
I cannot tell if the .d12 file has be attached to my previous post. I have therefore provided a image of my input file.
many thanks.


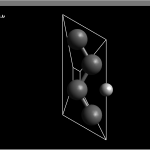

I am an undergraduate currently performing ab initio calculations to tunnel proton through a graphene supercell using CRYSTAL14
I have managed to tunnel hydrogen through, however upon changing the basis set of the hydrogen atom to a proton using CHEMOD i am not getting physical results.
Could you please advise me or direct me to someone who can help.